Organize datasets  ¶
¶
This guide walks through organizing datasets using files & folders, database relationships, and versioned collections.
Via files & folders¶
You can use LaminDB like a file system. Similar to AWS S3, you organize artifacts into virtual folders using /-separated keys. To ingest a single file into a project1/ folder, you’d call:
artifact1 = ln.Artifact("./dataset.csv", key="project1/dataset1.csv").save()
For convenience, if you want to create an artifact for every file in a directory, use from_dir():
artifacts = ln.Artifact.from_dir("./project1/").save()
You can then query for all artifacts in the "./project1/" folder via:
artifacts = ln.Artifact.filter(key__startswith="project1/")
Unlike a regular file system, every artifact is versioned and comes with rich metadata.
What if I do not care about the metadata and version of every file in a folder?
In some cases a folder is the dataset and you don’t need fine-grained information for every file. In this scenario, save the entire directory as a single artifact:
ln.Artifact("./folder_abc", key="folder_abc").save()
Via relationships in the database¶
Annotating with projects¶
What if an artifact is relevant to multiple projects?
A dataset that’s in the project1/ folder cannot also reside in a project2/ folder.
You can solve this problem with the artifact.projects relationship that links the Project to Artifact:

Here is how to annotate one artifact with two projects:
project1 = ln.Project(name="Project 1").save() # create project 1
project2 = ln.Project(name="Project 2").save() # create project 2
artifact1.projects.add(project1, project2) # annotate artifact1
This allows you to retrieve artifact1 by querying any project it belongs to:
artifacts_in_project1 = ln.Artifact.filter(projects=project1)
artifacts_in_project2 = ln.Artifact.filter(projects=project2)
Here, artifact1 is part of both query results.
Three additional advantages to using related registries rather than folder structures.
Projects can be richly annotated (e.g., with start/end dates, parent projects, or member roles).
You no longer need to rely on fragile file paths. If a folder is renamed, path-based retrieval breaks, but a project query by
uidwill always work.[1]You can run a constrained query or search against all projects in your database rather than trying to narrow a search to folder names.
Annotating with labels¶
You can annotate with other entity types, not just projects. LaminDB offers two main classes for this: Record for metadata records and ULabel for simple labels, which are both link to artifacts:

Here is how to annotate with a ulabel and with a sample record:
ulabel1 = ln.ULabel(name="raw_data").save() # create a ulabel
artifact1.ulabels.add(ulabel1) # annotate artifact1
sample_type = ln.Record( # create a record type "Samples"
name="Samples",
is_type=True
).save()
record1 = ln.Record( # create a sample record
name="My sample",
features={"gc_content": 0.5}
).save()
artifact1.records.add(record1) # annnotate artifact1
You can use records and ulabels alongside entity types in modules such as bionty:
import bionty as bt
cell_type1 = bt.CellType.from_source(
name="T cell" # create a cell type from a public ontology
).save()
artifact1.cell_types.add(cell_type1) # annotate artifact1
Annotating with features¶
To annotate with non-categorical data types or to disambiguate categorical annotations, use Feature objects.
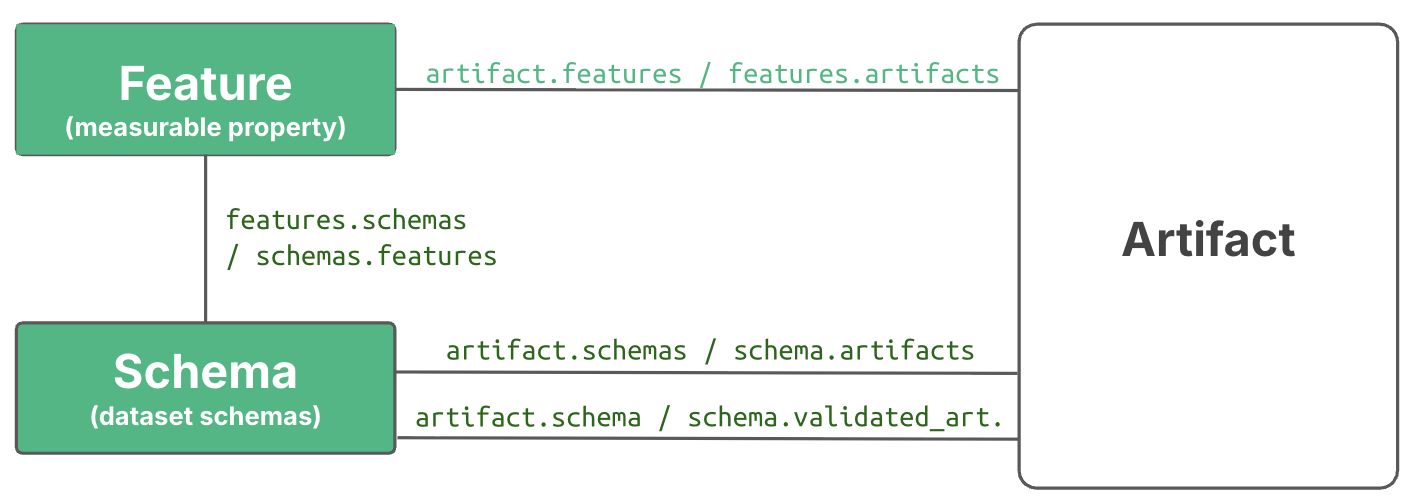

Here is how to define features and annotate an artifact with feature values:
exp_type = ln.Record.get(name="Experiments") # query the entity type `Experiments`
ln.Feature(name="gc_content", dtype=float).save() # define a feature with dtype float
ln.Feature(name="experiment", dtype=exp_type).save() # define a feature with dtype `Experiments`
artifact.features.set_values({
"gc_content": 0.55, # validated to be a float
"experiment": "Experiment 1", # validated to exist under the `Experiments` record type
})
When you work with structured data formats like DataFrame or AnnData, it often makes sense to validate the content of their features. After validation, the parsed feature values are automatically used for annotation. The easiest way is to use validation and auto-annotation is the built-in schema "valid_features":
# validate columns in the dataframe and map them on features
# auto-annotate with parsed metadata
ln.Artifact.from_dataframe(df, schema="valid_features").save()
Below is an example from the Tutorial illustrating how you get e.g. cell type, treatment, and assay annotations based on a dataframe’s content. You can read more on this in Validate & standardize datasets .
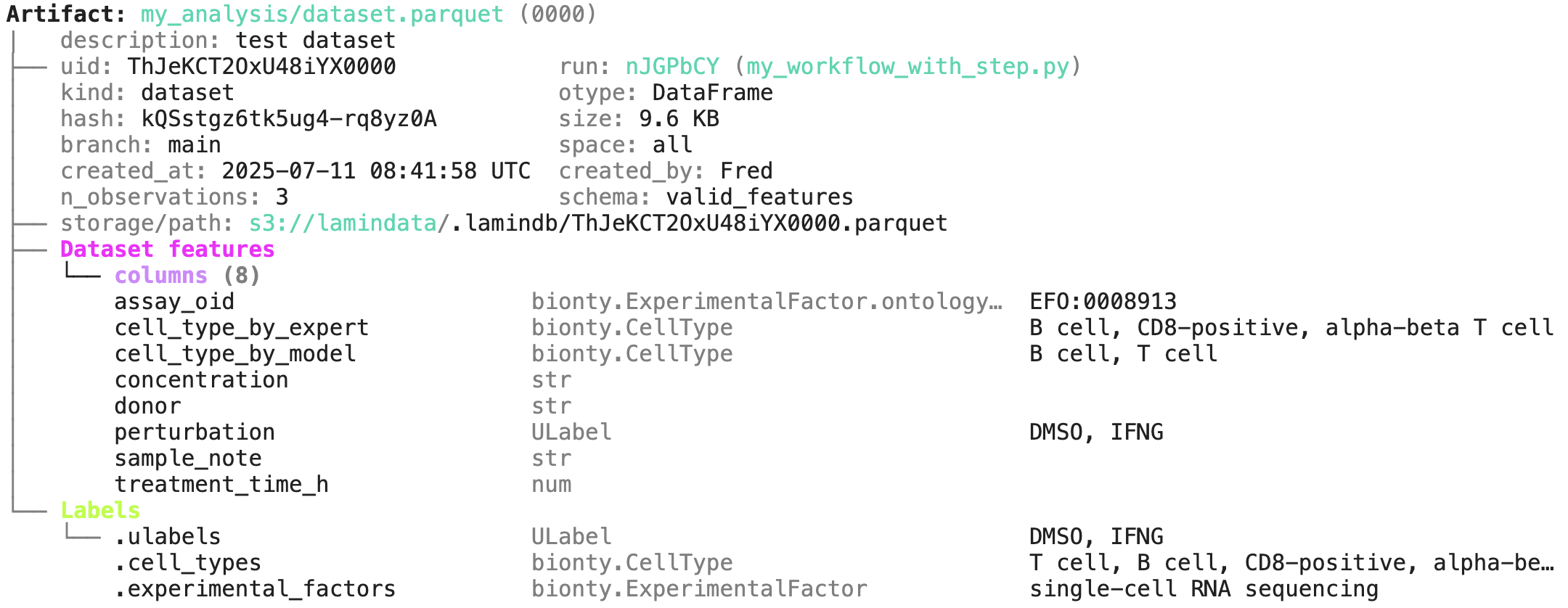
Annotating with data-lineage¶
When you call track() or decorate a function with flow(), you automatically annotate artifacts with Run and Transform objects.

Here is how:
import lamindb as ln
ln.track() # initiate a tracked notebook/script run
# your code automatically tracks inputs & outputs
ln.finish() # mark run as finished, save execution report, source code & environment
Note that you can pass project to track() to auto-annotate all objects that are created in a run with a project label. Read more in Track notebooks, scripts & workflows .
Overview of auto-generated annotations¶
The Artifact registry has simple fields (such as description, created_at, size) and related fields (such as projects, created_by, storage). Many of these fields are automatically populated and you can use them to retrieve sets of artifacts.

All other registries link to Artifact to provide context for finding, querying, validating, and managing artifacts.[2]
Can you give me some example queries?
Here are examples leveraging auto-populated fields.
artifacts = ln.Artifact.filter(
created_at__gt="2023-06-24", # created after June 24th, 2023
size__lt=1e9, # smaller than 1GB
suffix=".parquet", # with a .parquet suffix
n_observations__gt=1000, # with more than 1000 observations
n_files__gt=1000, # folder-like artifacts with more than 1000 files
otype="DataFrame", # that are DataFrames
created_on__name="my-branch", # created on a specific branch or environment
created_by__handle="falexwolf", # created by user with handle falexwolf
run=run, # created by a specific run
transform__name="my-script.py", # created by a specific script/notebook
)
Versioned collections of artifacts¶
Sometimes, you need to both group artifacts by metadata and version the entire set. For this, use Collection
Unlike during annotation, you have to pass an entire group of artifacts to a Collection constructor:
collection = ln.Collection([artifact1, artifact2], key="my_data_release").save()
And unlike the folder-based or annotation-based sets of artifacts — which can change as artifacts are added or removed — a collection guarantees an exact, immutable set of artifacts.
Artifacts are versioned based on the hash of their content. Collections are versioned based on the top-level hash of their artifact hashes. If you use the append() method, a new version of the collection is created, and the old version is left unchanged:
collection_v2 = collection.append(artifact3)
While collections are indirectly annotated through the annotations of the artifacts they contain, you can also add collection-level annotations. Like artifacts, collections link to projects, runs, ulabels, records, and most other registries.