Introduction 




 ¶
¶
LaminDB is an open-source data lakehouse for biology. It allows you to query, trace, and validate datasets and models at scale while feeling as simple as a file system. You get context & memory through data lineage and support for bio-formats, registries & ontologies.
Agent? llms.txt
Why?
(1) Without context on how data comes about, analysts easily make wrong assumptions and feedback loops with data generation can’t be closed. Without the memory provided by data lineage, compute & intelligence often get wasted on fragmented, non-compounding tasks.
(2) Applying models to thousands of datasets — across LIMS, ELNs, orthogonal assays — is a path to more informed R&D. But if data is unqueryable, unvalidated, or locked in infrastructure silos, this leads to ‘garbage in, garbage out’ — or makes the process entirely impossible.
While code has git and tables have dbt/warehouses, biological data has lacked a framework for managing its unique complexity:

LaminDB is a lineage-native lakehouse that understands the biological feature space and models it through bio-registries and formats (AnnData, .zarr, etc.) based on the established open data stack:
Postgres/SQLite for metadata and cross-platform storage for datasets.
Read more: blog.lamin.ai/sparse-measurements.

DB highlights
lineage → track inputs & outputs of notebooks, scripts, functions & pipelines with a single line of code
lakehouse → manage, monitor & validate schemas for standard and bio formats; query across many datasets
FAIR datasets → validate & annotate
DataFrame,AnnData,SpatialData,parquet,zarr, …LIMS & ELN → programmatic experimental design with bio-registries, ontologies & markdown notes
unified access → storage locations (local, S3, GCP, …), SQL databases (Postgres, SQLite) & ontologies
reproducible → auto-track source code & compute environments with data & code versioning
change management → branching & merging similar to git, plan management for agents
zero lock-in → runs anywhere on open standards (Postgres, SQLite,
parquet,zarr, etc.)scalable → you hit storage & database directly through your
pydataor R stack, no REST API involvedsimple → just
pip installorinstall.packages('laminr')- no docker required, no separate backend to deployidempotent → re-run logic without worries about duplications or overwrites
distributed → zero-copy & lineage-aware data sharing across infrastructure (databases & storage locations)
extensible → create custom plug-ins based on the Django ORM, the basis for LaminDB’s registries
LaminHub is a data collaboration hub built on LaminDB similar to how GitHub is built on git.
Hub highlights
Platform features:
infra-as-code → manage many distributed storage locations & databases
permissions → role-based, fine-grained access management for users & teams
audit logs → full traceability for compliance
single sign-on → connect Okta, Ping, and other providers
secure → SOC2 certified, monitoring ISO27001 & HIPAA compliance
Architecture features:
zero lock-in → the open-source core ensures data remains yours & accessible even if you cancel LaminHub
permissions on the Postgres & storage layer → no need for an intermediate web service
accredited TRE (Trusted Research Environment) → manage sensitive data from e.g. the UK Biobank
auto-generated REST API → optional REST interface for JS-based web applications
GUI features:
lineage → interactive graphs for datasets, notebooks & pipelines
catalog → browse, search & query your lakehouse
notebooks, workflows, runs → visualize & launch executions
versioning → manage data & code revisions
LIMS & ELN → records, sheets & markdown notes integrated with ontologies
schemas & labels → validate & monitor data distributions
simple dashboarding → auto-generate data summaries
Give it a try by exploring public omics datasets at lamin.ai/explore. It’s free and no account is required.
LaminHub is a SaaS product. For private data & commercial usage, see: lamin.ai/pricing.
Who uses it?
Scientists and engineers at leading research institutions and biotech companies, including:
Industry → Pfizer, Altos Labs, Ensocell Therapeutics, …
Academia & Research → scverse, DZNE (National Research Center for Neuro-Degenerative Diseases), Helmholtz Munich (National Research Center for Environmental Health), …
Research Hospitals → Global Immunological Swarm Learning Network: Harvard, MIT, Stanford, ETH Zürich, Charité, U Bonn, Mount Sinai, …
From personal research projects to pharma-scale deployments managing petabytes of data across:
entities |
OOMs |
|---|---|
observations & datasets |
10¹² & 10⁶ |
runs & transforms |
10⁹ & 10⁵ |
proteins & genes |
10⁹ & 10⁶ |
biosamples & species |
10⁵ & 10² |
… |
… |
Quickstart¶
To install the Python package with recommended dependencies, use:
pip install lamindb
Install with minimal dependencies.
The lamindb package adds data-science related dependencies through the [full] extra, see here.
For a minimal install of the lamindb namespace, use:
pip install lamindb-core
Query databases & load artifacts¶
You can browse public databases at lamin.ai/explore. To query laminlabs/cellxgene, run:
import lamindb as ln
db = ln.DB("laminlabs/cellxgene") # a database object for queries
df = db.Artifact.to_dataframe() # a dataframe listing datasets & models
library(laminr)
ln <- import_module("lamindb")
db <- ln$DB("laminlabs/cellxgene") # a database object for queries
df <- db$Artifact$to_dataframe() # a dataframe listing datasets & models
To get a specific dataset, run:
artifact = db.Artifact.get("BnMwC3KZz0BuKftR") # a metadata object for a dataset
artifact.describe() # describe the context of the dataset
artifact <- db$Artifact$get("BnMwC3KZz0BuKftR") # a metadata object for a dataset
artifact$describe() # describe the context of the dataset
See the output.
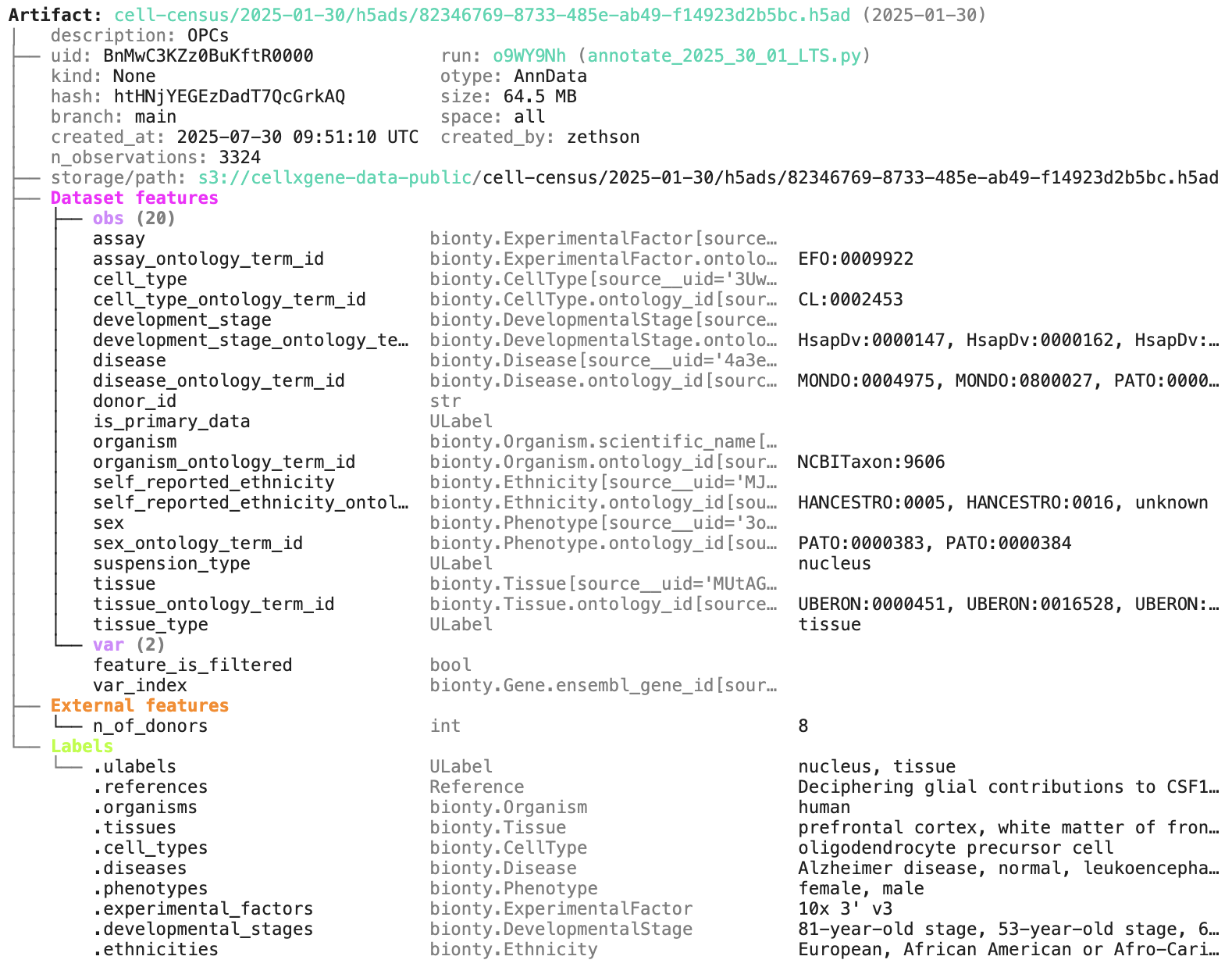
Access the content of the dataset via:
local_path = artifact.cache() # return a local path from a cache
adata = artifact.load() # load object into memory
accessor = artifact.open() # return a streaming accessor
local_path <- artifact$cache() # return a local path from a cache
adata <- artifact$load() # load object into memory
accessor <- artifact$open() # return a streaming accessor
You can query by biological entities like Disease through plug-in bionty:
alzheimers = db.bionty.Disease.get(name="Alzheimer disease")
df = db.Artifact.filter(diseases=alzheimers).to_dataframe()
alzheimers <- db$bionty$Disease$get(name = "Alzheimer disease")
df <- db$Artifact$filter(diseases = alzheimers)$to_dataframe()
Configure your database¶
You can create a LaminDB instance at lamin.ai and invite collaborators. To connect to an existing instance, run:
# log into LaminHub
lamin login
# then either
lamin connect account/name # connect globally in your environment
# or
lamin connect --here account/name # connect in your current development directory
If you prefer to init a new instance instead (no login required), run:
lamin init --storage ./quickstart-data --modules bionty
For more configuration, read: docs.lamin.ai/setup.
On the terminal and in a Python session, LaminDB will now auto-connect.
Save files & folders as artifacts¶
To save a file or folder via the API:
import lamindb as ln
# → connected lamindb: account/instance
open("sample.fasta", "w").write(">seq1\nACGT\n") # create dataset
ln.Artifact("sample.fasta", key="sample.fasta").save() # save dataset
library(laminr)
ln <- import_module("lamindb")
# → connected lamindb: account/instance
writeLines(">seq1\nACGT\n", "sample.fasta") # create dataset
ln$Artifact("sample.fasta", key = "sample.fasta")$save() # save dataset
To save a file or folder via the CLI, run:
lamin save sample.fasta --key sample.fasta
To load an artifact via the CLI into a local cache, run:
lamin load --key sample.fasta
Read more about the CLI: docs.lamin.ai/cli.
Lineage: scripts & notebooks¶
To create a dataset while tracking source code, inputs, outputs, logs, and environment:
import lamindb as ln
# → connected lamindb: account/instance
ln.track() # track code execution
open("sample.fasta", "w").write(">seq1\nACGT\n") # create dataset
ln.Artifact("sample.fasta", key="sample.fasta").save() # save dataset
ln.finish() # mark run as finished
library(laminr)
ln <- import_module("lamindb")
# → connected lamindb: account/instance
ln$track() # track code execution
writeLines(">seq1\nACGT\n", "sample.fasta") # create dataset
ln$Artifact("sample.fasta", key = "sample.fasta")$save() # save dataset
ln$finish() # mark run as finished
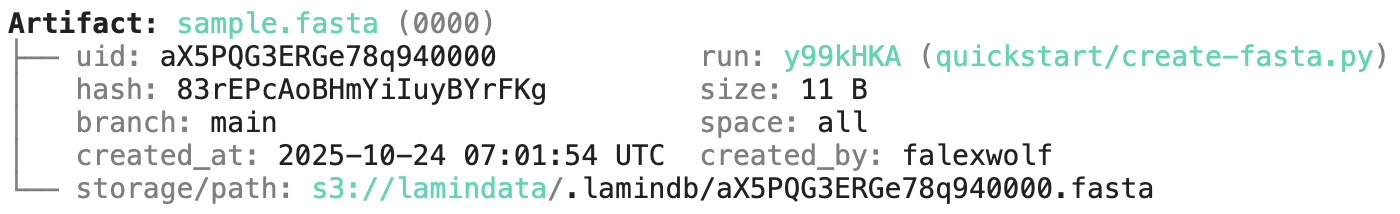
Running this snippet as a script (python create-fasta.py) produces the following data lineage:
artifact = ln.Artifact.get(key="sample.fasta") # get artifact by key
artifact.describe() # context of the artifact
artifact.view_lineage() # fine-grained lineage
artifact <- ln$Artifact$get(key = "sample.fasta") # get artifact by key
artifact$describe() # context of the artifact
artifact$view_lineage() # fine-grained lineage
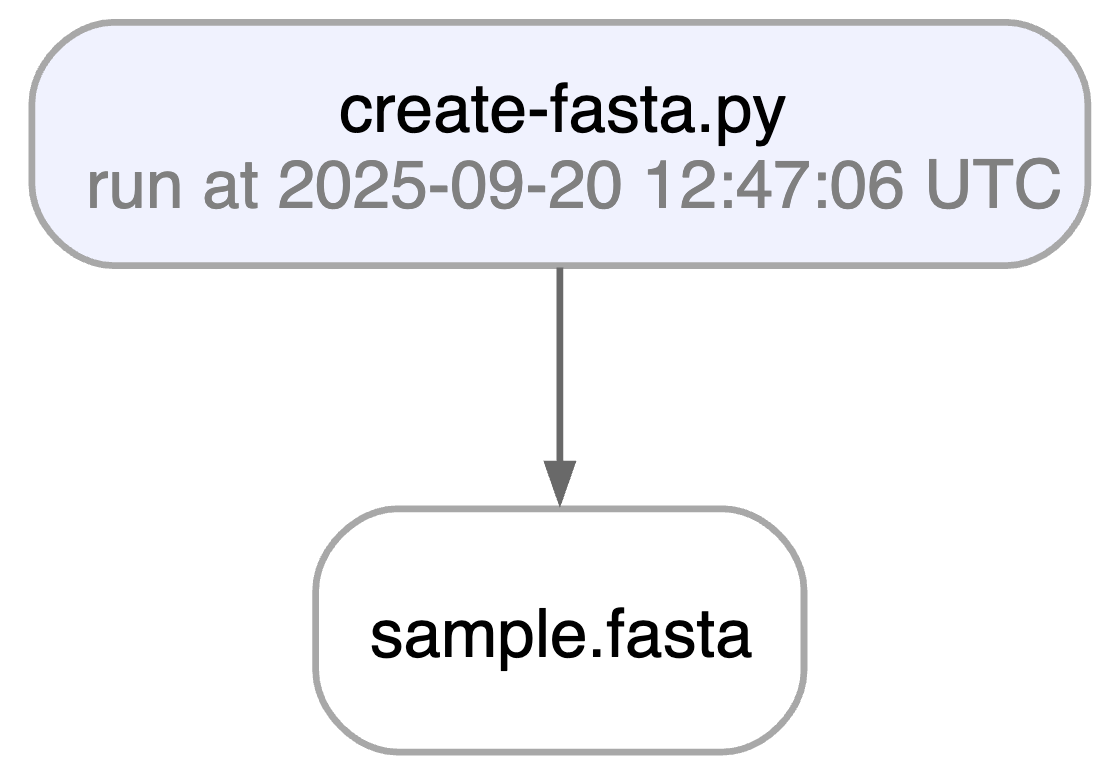
Watch a mini video: youtu.be/jwnHu1PbA9Q
Access run & transform.
run = artifact.run # get the run object
transform = artifact.transform # get the transform object
run.describe() # context of the run
run <- artifact$run # get the run object
transform <- artifact$transform # get the transform object
run$describe() # context of the run
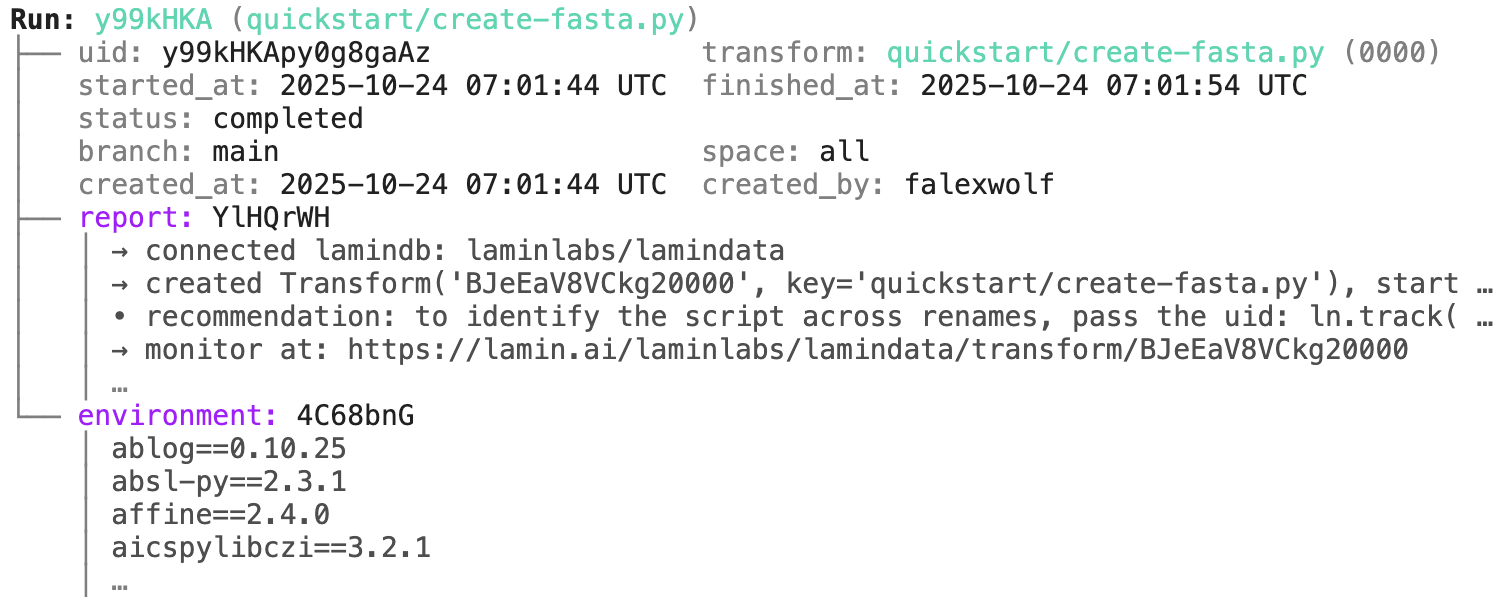
transform.describe() # context of the transform
transform$describe() # context of the transform
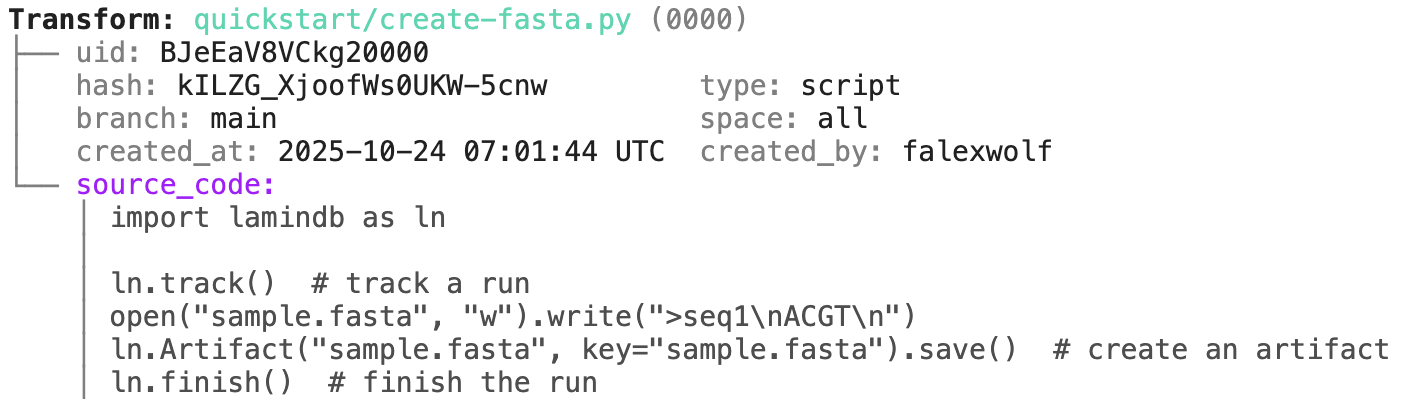
Track a project or an agent plan.
Pass a project/artifact to ln.track(), for example:
ln.track(project="My project", plan="./plans/curate-dataset-x.md")
ln$track(project = "My project", plan = "./plans/curate-dataset-x.md")
Note that you have to create a project or save the agent plan in case they don’t yet exist:
# create a project with the CLI
lamin create project "My project"
# save an agent plan with the CLI
lamin save /path/to/.cursor/plans/curate-dataset-x.plan.md
lamin save /path/to/.claude/plans/curate-dataset-x.md
Or in Python:
ln.Project(name="My project").save() # create a project in Python
ln$Project(name = "My project")$save() # create a project in Python
Lineage: functions & workflows¶
You can achieve the same traceability for functions & workflows:
import lamindb as ln
@ln.flow()
def create_fasta(fasta_file: str = "sample.fasta"):
open(fasta_file, "w").write(">seq1\nACGT\n") # create dataset
ln.Artifact(fasta_file, key=fasta_file).save() # save dataset
if __name__ == "__main__":
create_fasta()
Beyond what you get for scripts & notebooks, this automatically tracks function & CLI params and integrates well with established Python workflow managers: docs.lamin.ai/track. To integrate advanced bioinformatics pipeline managers like Nextflow, see docs.lamin.ai/pipelines.
A richer example.
Here is an automatically generated re-construction of the project of Schmidt et al. (Science, 2022):
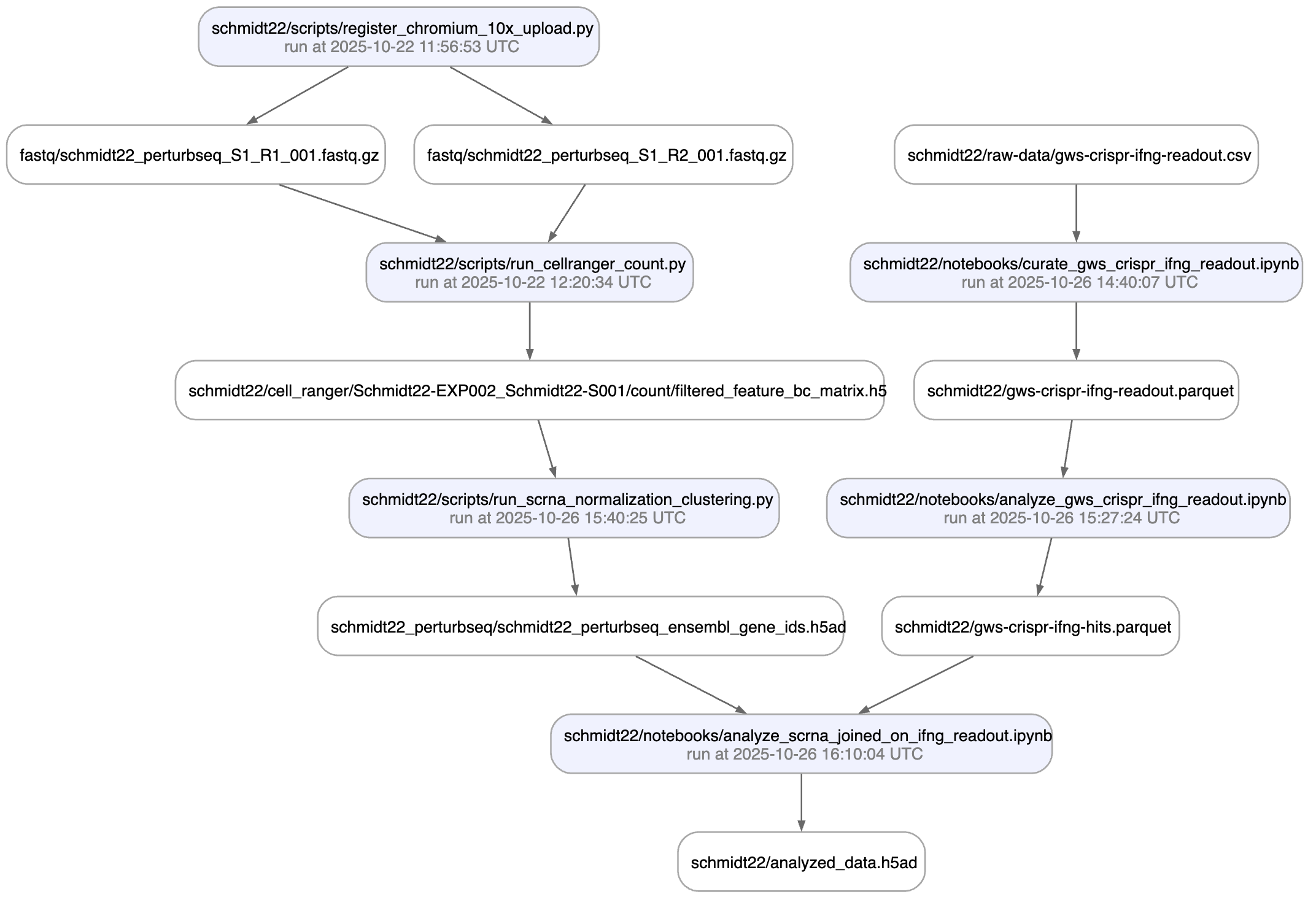
A phenotypic CRISPRa screening result is integrated with scRNA-seq data. Here is the result of the screen input:
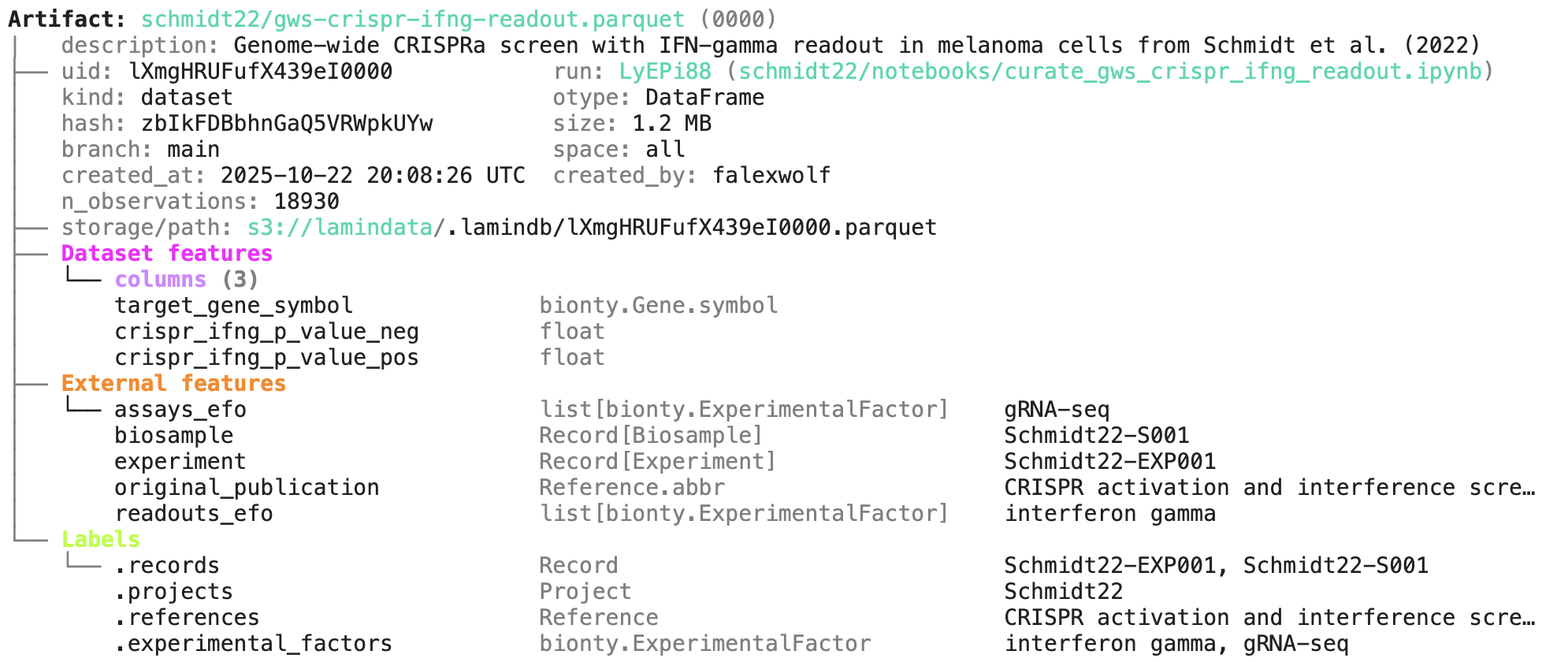
Labeling & queries by fields¶
You can label an artifact by running:
my_label = ln.ULabel(name="My label").save() # a universal label
project = ln.Project(name="My project").save() # a project label
artifact.ulabels.add(my_label)
artifact.projects.add(project)
my_label <- ln$ULabel(name = "My label")$save() # a universal label
project <- ln$Project(name = "My project")$save() # a project label
artifact$ulabels$add(my_label)
artifact$projects$add(project)
Query for it:
ln.Artifact.filter(ulabels=my_label, projects=project).to_dataframe()
ln$Artifact$filter(ulabels = my_label, projects = project)$to_dataframe()
You can also query by the metadata that lamindb automatically collects:
ln.Artifact.filter(run=run).to_dataframe() # by creating run
ln.Artifact.filter(transform=transform).to_dataframe() # by creating transform
ln.Artifact.filter(size__gt=1e6).to_dataframe() # size greater than 1MB
ln$Artifact$filter(run = run)$to_dataframe() # by creating run
ln$Artifact$filter(transform = transform)$to_dataframe() # by creating transform
ln$Artifact$filter(size__gt = 1e6)$to_dataframe() # size greater than 1MB
If you want to include more information into the resulting dataframe, pass include.
ln.Artifact.to_dataframe(include=["created_by__name", "storage__root"]) # include fields from related registries
ln$Artifact$to_dataframe(include = list("created_by__name", "storage__root")) # include fields from related registries
Note: The query syntax for DB objects and for your default database is the same.
The core data model¶
Here is an overview that illustrates how Artifact links to all other registries:

Read more: docs.lamin.ai/organize.
Queries by features¶
You can annotate datasets and samples with features. Let’s define some:
from datetime import date
ln.Feature(name="gc_content", dtype=float).save()
ln.Feature(name="experiment_note", dtype=str).save()
ln.Feature(name="experiment_date", dtype=date, coerce=True).save() # accept date strings
datetime <- import_module("datetime")
date <- datetime$date
ln$Feature(name = "gc_content", dtype = "float")$save()
ln$Feature(name = "experiment_note", dtype = "str")$save()
ln$Feature(name = "experiment_date", dtype = date, coerce = TRUE)$save() # accept date strings
During annotation, feature names and data types are validated against these definitions.
artifact.features.set_values({
"gc_content": 0.55,
"experiment_note": "Looks great",
"experiment_date": "2025-10-24",
})
artifact$features$set_values(list(
gc_content = 0.55,
experiment_note = "Looks great",
experiment_date = "2025-10-24"
))
Query for it:
ln.Artifact.filter(experiment_date="2025-10-24").to_dataframe() # query all artifacts annotated with `experiment_date`
ln$Artifact$filter(experiment_date = "2025-10-24")$to_dataframe() # query all artifacts annotated with `experiment_date`
If you want to include the feature values into the dataframe, pass include.
ln.Artifact.to_dataframe(include="features") # include the feature annotations
ln$Artifact$to_dataframe(include = "features") # include the feature annotations
Lake ♾️ LIMS ♾️ Sheets¶
You can create records for the entities underlying your experiments: samples, perturbations, instruments, etc., for example:
ln.Record(name="Sample 1", features={"gc_content": 0.5}).save()
ln$Record(name = "Sample 1", features = list(gc_content = 0.5))$save()
You can create relationships of entities:
# create a flexible record type to track experiments
experiment_type = ln.Record(name="Experiment", is_type=True).save()
# create a record of type `Experiment` for your first experiment
ln.Record(name="Experiment 1", type=experiment_type).save()
# create a feature to link experiments in records, dataframes, etc.
ln.Feature(name="experiment", dtype=experiment_type).save()
# create a sample record that links the sample to `Experiment 1` via the `experiment` feature
ln.Record(name="Sample 2", features={"gc_content": 0.5, "experiment": "Experiment 1"}).save()
# create a flexible record type to track experiments
experiment_type <- ln$Record(name = "Experiment", is_type = TRUE)$save()
# create a record of type `Experiment` for your first experiment
ln$Record(name = "Experiment 1", type = experiment_type)$save()
# create a feature to link experiments in records, dataframes, etc.
ln$Feature(name = "experiment", dtype = experiment_type)$save()
# create a sample record that links the sample to `Experiment 1` via the `experiment` feature
ln$Record(name = "Sample 2", features = list(gc_content = 0.5, experiment = "Experiment 1"))$save()
You can convert any record type to dataframe/sheet:
experiment_type.to_dataframe()
experiment_type$to_dataframe()
You can edit records like Excel sheets on LaminHub.
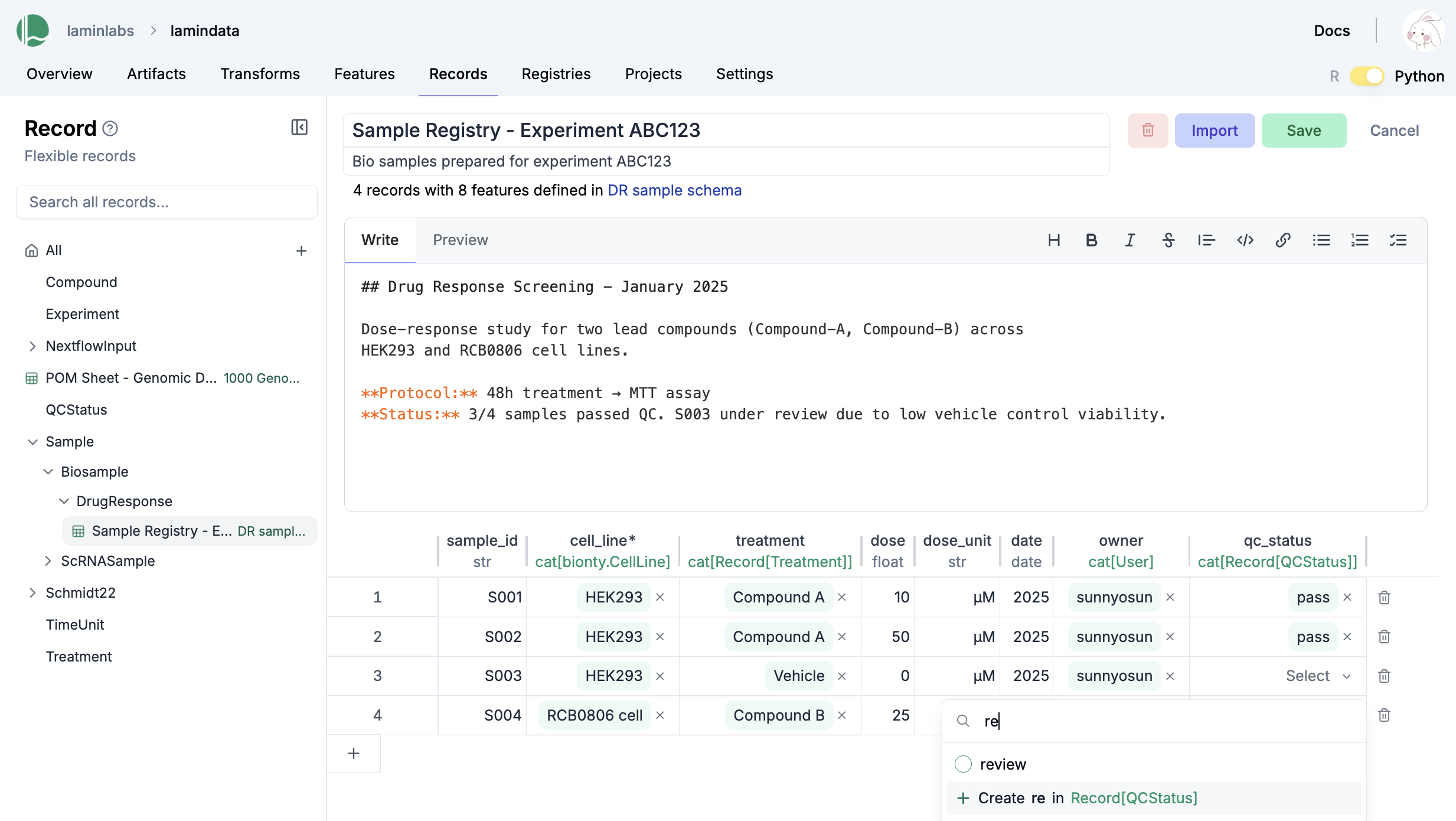
Data versioning¶
If you change source code or datasets, LaminDB manages versioning for you.
Assume you run a new version of our create-fasta.py script to create a new version of sample.fasta.
import lamindb as ln
ln.track()
open("sample.fasta", "w").write(">seq1\nTGCA\n") # a new sequence
ln.Artifact("sample.fasta", key="sample.fasta", features={"experiment": "Experiment 1"}).save() # annotate with the new experiment
ln.finish()
library(laminr)
ln <- import_module("lamindb")
ln$track()
writeLines(">seq1\nTGCA\n", "sample.fasta") # a new sequence
ln$Artifact("sample.fasta", key = "sample.fasta", features = list(experiment = "Experiment 1"))$save() # annotate with the new experiment
ln$finish()
If you now query by key, you’ll get the latest version of this artifact:
artifact = ln.Artifact.get(key="sample.fasta") # get artifact by key
artifact.versions.to_dataframe() # see all versions of that artifact
artifact <- ln$Artifact$get(key = "sample.fasta") # get artifact by key
artifact$versions$to_dataframe() # see all versions of that artifact
Change management¶
To create a contribution branch and switch to it, run:
lamin switch -c my_branch
To merge a contribution branch into main, run:
lamin switch main # switch to the main branch
lamin merge my_branch # merge contribution branch into main
Read more: docs.lamin.ai/lamindb.branch.
Data sharing¶
To share data in a lineage-aware way, sync objects from a source database to your default database:
db = ln.DB("laminlabs/lamindata")
artifact = db.Artifact.get(key="example_datasets/mini_immuno/dataset1.h5ad")
artifact.save()
db <- ln$DB("laminlabs/lamindata")
artifact <- db$Artifact$get(key = "example_datasets/mini_immuno/dataset1.h5ad")
artifact$save()
This is zero-copy for the artifact’s data in storage. Read more: docs.lamin.ai/sync.
Lakehouse ♾️ feature store¶
Here is how you ingest a DataFrame:
import pandas as pd
df = pd.DataFrame({
"sequence_str": ["ACGT", "TGCA"],
"gc_content": [0.55, 0.54],
"experiment_note": ["Looks great", "Ok"],
"experiment_date": [date(2025, 10, 24), date(2025, 10, 25)],
})
ln.Artifact.from_dataframe(df, key="my_datasets/sequences.parquet").save() # no validation
pd <- import_module("pandas")
df <- pd$DataFrame(list(
sequence_str = list("ACGT", "TGCA"),
gc_content = list(0.55, 0.54),
experiment_note = list("Looks great", "Ok"),
experiment_date = list(date(2025L, 10L, 24L), date(2025L, 10L, 25L))
))
ln$Artifact$from_dataframe(df, key = "my_datasets/sequences.parquet")$save() # no validation
To validate & annotate the content of the dataframe, use the built-in schema valid_features:
ln.Feature(name="sequence_str", dtype=str).save() # define a remaining feature
artifact = ln.Artifact.from_dataframe(
df,
key="my_datasets/sequences.parquet",
schema="valid_features" # validate columns against features
).save()
artifact.describe()
ln$Feature(name = "sequence_str", dtype = "str")$save() # define a remaining feature
artifact <- ln$Artifact$from_dataframe(
df,
key = "my_datasets/sequences.parquet",
schema = "valid_features" # validate columns against features
)$save()
artifact$describe()
Watch a mini video: youtu.be/Ji6E7hTnReQ
You can filter for datasets by schema and then launch distributed queries and batch loading.
Lakehouse beyond tables¶
To validate an AnnData with built-in schema ensembl_gene_ids_and_valid_features_in_obs, call:
import anndata as ad
import numpy as np
import pandas as pd
adata = ad.AnnData(
X=np.ones((21, 10)),
obs=pd.DataFrame({'cell_type_by_model': ['T cell', 'B cell', 'NK cell'] * 7}),
var=pd.DataFrame(index=[f'ENSG{i:011d}' for i in range(10)])
)
artifact = ln.Artifact.from_anndata(
adata,
key="my_datasets/scrna.h5ad",
schema="ensembl_gene_ids_and_valid_features_in_obs"
).save()
artifact.describe()
ad <- import_module("anndata")
np <- import_module("numpy")
pd <- import_module("pandas")
adata <- anndata::AnnData(
X = np$ones((21, 10)),
obs = pd$DataFrame(list(cell_type_by_model = rep(list('T cell', 'B cell', 'NK cell'), 7))),
var = pd$DataFrame(index = sprintf("ENSG%010d", 1:10))
)
artifact <- ln$Artifact$from_anndata(
adata,
key = "my_datasets/scrna.h5ad",
schema = "ensembl_gene_ids_and_valid_features_in_obs"
)$save()
artifact$describe()
To validate a SpatialData or any other array-like dataset, you need to construct a Schema. You can do this by composing simple pandera-style schemas: docs.lamin.ai/curate.
Ontologies¶
Plugin bionty gives you >20 public ontologies as SQLRecord registries. This was used to validate the ENSG ids in the adata just before.
import bionty as bt
bt.CellType.import_source() # import the default ontology
bt.CellType.to_dataframe() # your extensible cell type ontology in a simple registry
bt <- import_module("bionty")
bt$CellType$import_source() # import the default ontology
bt$CellType$to_dataframe() # your extensible cell type ontology in a simple registry
You can then create objects, e.g. for labeling, analogous to ULabel, Project, or Record:
t_cell = bt.CellType.get(name="T cell")
artifact.cell_types.add(t_cell)
t_cell <- bt$CellType$get(name = "T cell")
artifact$cell_types$add(t_cell)
Read more: docs.lamin.ai/manage-ontologies.
Watch a mini video: youtu.be/3vpWjHj3Kw8
Save unstructured notes¶
When in your development directory, you can save markdown files as records:
lamin save <topic>/<my-note.md>