Append a new dataset 
 ¶
¶
We have one dataset in storage and are about to receive a new dataset.
In this notebook, we’ll see how to manage the situation.
import lamindb as ln
import bionty as bt
import readfcs
bt.settings.organism = "human"
ln.track("SmQmhrhigFPL0000")
Show code cell output
→ connected lamindb: testuser1/test-facs
→ created Transform('SmQmhrhigFPL0000', key='facs2.ipynb'), started new Run('L5HBYiJIpQdoAozc') at 2026-05-20 14:53:51 UTC
→ notebook imports: bionty==2.4.0 lamindb-core==2.4.2 pytometry==0.1.6 readfcs==2.1.0 scanpy==1.12.1
Ingest a new artifact¶
Access  ¶
¶
Let us validate and register another .fcs file from Oetjen18:
filepath = readfcs.datasets.Oetjen18_t1()
adata = readfcs.read(filepath)
# since anndata>=0.12.0, `/` is not allowed in keys
adata.var.index = adata.var.index.str.replace("/", "|")
adata.var["marker"] = adata.var["marker"].str.replace("/", "|")
adata.uns["meta"]["spill"].index = adata.uns["meta"]["spill"].index.str.replace(
"/", "|"
)
adata.uns["meta"]["spill"].columns = adata.uns["meta"]["spill"].columns.str.replace(
"/", "|"
)
adata
Show code cell output
/opt/hostedtoolcache/Python/3.12.13/x64/lib/python3.12/functools.py:912: ImplicitModificationWarning: Transforming to str index.
return dispatch(args[0].__class__)(*args, **kw)
AnnData object with n_obs × n_vars = 241552 × 20
var: 'n', 'channel', 'marker', 'PnR', 'PnB', 'PnE', 'PnV', 'PnG'
uns: 'meta'
Transform: normalize  ¶
¶
import pytometry as pm
pm.pp.split_signal(adata, var_key="channel")
Show code cell output
/opt/hostedtoolcache/Python/3.12.13/x64/lib/python3.12/site-packages/pytometry/pp/_process_data.py:231: FutureWarning: Use var (e.g. `k in adata.var` or `str(adata.var.columns.tolist())`) instead of AnnData.var_keys, AnnData.var_keys is deprecated and will be removed in the future.
if key_in not in adata.var_keys():
/opt/hostedtoolcache/Python/3.12.13/x64/lib/python3.12/site-packages/pytometry/pp/_process_data.py:65: FutureWarning: Use var (e.g. `k in adata.var` or `str(adata.var.columns.tolist())`) instead of AnnData.var_keys, AnnData.var_keys is deprecated and will be removed in the future.
elif var_key in adata.var_keys():
pm.pp.compensate(adata)
pm.tl.normalize_biExp(adata)
adata = adata[ # subset to rows that do not have nan values
adata.to_df().isna().sum(axis=1) == 0
]
adata.to_df().describe()
Show code cell output
| CD95 | CD8 | CD27 | CXCR4 | CCR7 | LIVE|DEAD | CD4 | CD45RA | CD3 | CD49B | CD14|19 | CD69 | CD103 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| count | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 | 241552.000000 |
| mean | 887.579860 | 1302.985717 | 1221.257257 | 877.533482 | 977.505533 | 1883.358298 | 556.687953 | 929.493316 | 941.166747 | 966.012244 | 1210.769935 | 741.523184 | 1003.064857 |
| std | 573.549695 | 827.850302 | 672.851319 | 411.966073 | 584.217139 | 932.113729 | 480.875917 | 795.550133 | 658.984751 | 456.437094 | 694.622980 | 473.287558 | 642.728024 |
| min | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
| 25% | 462.757715 | 493.413744 | 605.463427 | 588.047798 | 495.437303 | 1063.670965 | 240.623098 | 404.087640 | 477.932659 | 592.294399 | 575.401173 | 380.247262 | 475.108131 |
| 50% | 774.350833 | 1207.624048 | 1110.367681 | 782.939692 | 782.981430 | 1951.855099 | 484.355203 | 557.904360 | 655.909639 | 800.280049 | 1124.574275 | 705.802991 | 775.101973 |
| 75% | 1327.792103 | 2036.849496 | 1721.730010 | 1070.479036 | 1453.929567 | 2623.975657 | 729.754419 | 1345.771633 | 1218.445208 | 1347.042403 | 1742.288464 | 1069.175380 | 1420.744291 |
| max | 4053.903716 | 4065.495666 | 4095.351322 | 4025.827267 | 3999.075551 | 4096.000000 | 4088.719985 | 3961.255364 | 3940.061146 | 4089.445928 | 3982.769373 | 3810.774988 | 4023.968008 |
Validate cell markers  ¶
¶
Let’s see how many markers validate:
validated = bt.CellMarker.validate(adata.var.index)
Show code cell output
! 9 unique terms (69.20%) are not validated for name: 'CD95', 'CXCR4', 'CCR7', 'LIVE|DEAD', 'CD4', 'CD49B', 'CD14|19', 'CD69', 'CD103'
Let’s standardize and re-validate:
adata.var.index = bt.CellMarker.standardize(adata.var.index)
validated = bt.CellMarker.validate(adata.var.index)
Show code cell output
! 7 unique terms (53.80%) are not validated for name: 'CD95', 'CXCR4', 'LIVE|DEAD', 'CD49B', 'CD14|19', 'CD69', 'CD103'
/tmp/ipykernel_3613/92294437.py:1: ImplicitModificationWarning: Trying to modify index of attribute `.var` of view, initializing view as actual.
adata.var.index = bt.CellMarker.standardize(adata.var.index)
Next, register non-validated markers from Bionty:
records = bt.CellMarker.from_values(adata.var.index[~validated])
ln.save(records)
Manually create 1 marker:
bt.CellMarker(name="CD14|19").save()
Show code cell output
CellMarker(uid='67ZpJGSKNFyEAq', abbr=None, synonyms=None, description=None, name='CD14|19', gene_symbol=None, ncbi_gene_id=None, uniprotkb_id=None, branch_id=1, created_on_id=1, space_id=1, created_by_id=1, run_id=2, source_id=None, organism_id=1, created_at=2026-05-20 14:53:55 UTC, is_locked=False)
Move metadata to obs:
validated = bt.CellMarker.validate(adata.var.index)
adata.obs = adata[:, ~validated].to_df()
adata = adata[:, validated].copy()
Show code cell output
! 1 unique term (7.70%) is not validated for name: 'LIVE|DEAD'
Now all markers pass validation:
validated = bt.CellMarker.validate(adata.var.index)
assert all(validated)
Register  ¶
¶
schema = ln.Schema.get(name="FACS-AnnData-schema")
curator = ln.curators.AnnDataCurator(adata, schema=schema)
artifact = curator.save_artifact(description="Oetjen18_t1")
Show code cell output
! 1 term not validated in feature 'columns' in slot 'obs': 'LIVE|DEAD'
→ fix typos, remove non-existent values, or save terms via: curator.slots['obs'].cat.add_new_from('columns')
! no values were validated for columns!
→ returning schema with same hash: Schema(uid='0000000000000000', is_type=False, name='FACS-sample-metadata', description=None, n_members=None, coerce=None, flexible=True, itype='Feature', otype=None, hash='kMi7B_N88uu-YnbTLDU-DA', minimal_set=True, ordered_set=False, maximal_set=False, branch_id=1, created_on_id=1, space_id=1, created_by_id=1, run_id=1, type_id=None, created_at=2026-05-20 14:53:45 UTC, is_locked=False)
Annotate with more labels:
efs = bt.ExperimentalFactor.lookup()
organism = bt.Organism.lookup()
artifact.labels.add(efs.fluorescence_activated_cell_sorting)
artifact.labels.add(organism.human)
artifact.describe()
Show code cell output
Artifact: (0000) | description: Oetjen18_t1 ├── uid: 5ZHKrCmCU6KMWin60000 run: L5HBYiJ (facs2.ipynb) │ kind: dataset otype: AnnData │ hash: BkQOx3xp3OR4FoOq4CsuJA size: 44.4 MB │ branch: main space: all │ created_at: 2026-05-20 14:53:56 UTC created_by: testuser1 │ n_observations: 241552 schema: FACS-AnnData-schema ├── storage/path: /home/runner/work/lamin-usecases/lamin-usecases/docs/test-facs/.lamindb/5ZHKrCmCU6KMWin60000.h5ad ├── Dataset features │ ├── obs (None) │ └── var.T (12 bionty.CellMarker) │ CD103 num │ CD14|19 num │ CD27 num │ CD3 num │ CD45RA num │ CD49B num │ CD69 num │ CD8 num │ CD95 num │ CXCR4 num │ Ccr7 num │ Cd4 num └── Labels └── .organisms bionty.Organism human .experimental_factors bionty.ExperimentalFactor fluorescence-activated cell sorting
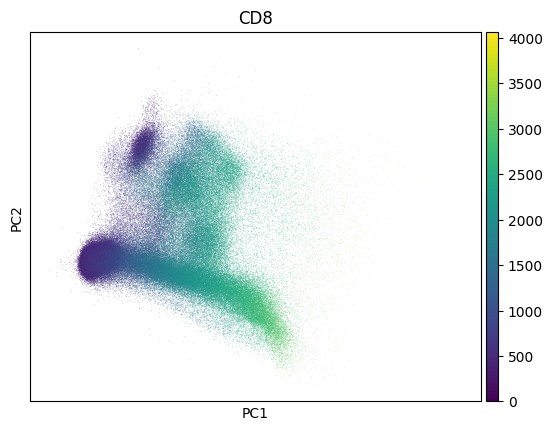
Inspect a PCA fo QC - this collection looks much like noise:
import scanpy as sc
markers = bt.CellMarker.lookup()
sc.pp.pca(adata)
sc.pl.pca(adata, color=markers.cd8.name)
Show code cell output
Create a new version of the collection by appending a artifact¶
Query the old version:
collection_v1 = ln.Collection.get(key="My versioned cytometry collection")
collection_v2 = ln.Collection(
[artifact, collection_v1.ordered_artifacts[0]],
revises=collection_v1,
version="2",
)
collection_v2.describe()
Show code cell output
Collection: My versioned cytometry collection (2) └── uid: iwtqnzhxu5uRsGnx0001 run: L5HBYiJ (facs2.ipynb) branch: main space: all created_at: <django.db.models.expressions.DatabaseDefault object at 0x7f5063152e70> created_by: testuser1
collection_v2.save()
Show code cell output
Collection(uid='iwtqnzhxu5uRsGnx0001', key='My versioned cytometry collection', description=None, hash='eNSQbRNsjg278G1RwXmqCw', reference=None, reference_type=None, meta_artifact=None, branch_id=1, created_on_id=1, space_id=1, created_by_id=1, run_id=2, created_at=2026-05-20 14:53:57 UTC, is_locked=False, version_tag='2', is_latest=True)