Vitessce: AnnData 
 ¶
¶
This tutorial demonstrates how to use Vitessce to create interactive visualizations for data stored as LaminDB artifacts. It requires a remote LaminDB instance with cloud storage to enable the Vitessce button (shown below) in the web interface.

It has been adapted from the data preparation examples in the Vitessce documentation.
In this part, we’ll visualize an AnnData object stored in both H5AD and Zarr formats.
# pip install 'vitessce[all]>=3.5.0' lamindb
!lamin connect laminlabs/lamindata # <-- replace with your remote instance
Show code cell output
→ connected lamindb: laminlabs/lamindata
• to map a local dev directory, call: lamin settings set dev-dir .
import vitessce as vit
import lamindb as ln
ln.track()
Show code cell output
→ connected lamindb: laminlabs/lamindata
→ found notebook vitessce.ipynb, making new version -- anticipating changes
→ created Transform('ZOf7HJEqB4y9000C', key='vitessce.ipynb'), started new Run('seATHQDWnvTX7QIM') at 2026-05-26 09:52:30 UTC
→ notebook imports: lamindb-core==2.4.2 vitessce==3.8.4
• recommendation: to identify the notebook across renames, pass the uid: ln.track("ZOf7HJEqB4y9")
Visualize an AnnData object (H5AD format)¶
Here we use the Habib et al. 2017 dataset from the COVID-19 Cell Atlas that has been previously subset to highly variable genes. The dataset was ingested into the public laminlabs/lamindata instance in this transform.
h5ad_artifact = ln.Artifact.get(key="vitessce_examples/habib17.h5ad")
When using .h5ad files, we construct a Reference Specification which enables interoperability with the Zarr interface.
The Reference Specification JSON was also generated in the transform above.
ref_artifact = ln.Artifact.get(key="vitessce_examples/habib17.reference.json")
Save a VitessceConfig object¶
You can create a dashboard for one or several datasets by using Vitessce’s component API.
You can pass artifacts to the AnnDataWrapper class using the adata_artifact and ref_artifact parameters.
vc = vit.VitessceConfig(
schema_version="1.0.18",
description=h5ad_artifact.description,
)
dataset = vc.add_dataset(name="Habib 2017").add_object(
vit.AnnDataWrapper(
adata_artifact=h5ad_artifact,
ref_artifact=ref_artifact,
obs_feature_matrix_path="X",
obs_embedding_paths=["obsm/X_umap"],
obs_embedding_names=["UMAP"],
obs_set_paths=["obs/CellType"],
obs_set_names=["Cell Type"],
)
)
obs_sets = vc.add_view(vit.ViewType.OBS_SETS, dataset=dataset)
obs_sets_sizes = vc.add_view(vit.ViewType.OBS_SET_SIZES, dataset=dataset)
scatterplot = vc.add_view(vit.ViewType.SCATTERPLOT, dataset=dataset, mapping="UMAP")
heatmap = vc.add_view(vit.ViewType.HEATMAP, dataset=dataset)
genes = vc.add_view(vit.ViewType.FEATURE_LIST, dataset=dataset)
vc.link_views([scatterplot, heatmap], ["featureValueColormapRange"], [[0.0, 0.1]])
vc.layout(((scatterplot | obs_sets) / heatmap) | (obs_sets_sizes / genes))
Show code cell output
<vitessce.config.VitessceConfig at 0x7fc06f5fe900>
Save the VitessceConfig object.
h5ad_vc_artifact = ln.integrations.save_vitessce_config(
vc,
description="View Habib17 (h5ad) in Vitessce",
)
Show code cell output
→ VitessceConfig references these artifacts:
Artifact(uid='NNUzTpWz3a0PU2aT0000', key='vitessce_examples/habib17.h5ad', description='Habib et al., 2017 Nature Methods, h5ad', suffix='.h5ad', kind='dataset', otype='AnnData', size=22586523, hash='nBetKKNFs4mjJ04X_f4uaA', n_files=None, n_observations=None, branch_id=1, created_on_id=1, space_id=1, storage_id=2, run_id=1932, schema_id=None, created_by_id=40, created_at=2026-01-20 19:52:45 UTC, is_locked=False, version_tag=None, is_latest=True)
Artifact(uid='cyzNRxkUQF1sMVrr0000', key='vitessce_examples/habib17.reference.json', description='Reference JSON for H5AD file, Habib et al., 2017 Nature Methods', suffix='.json', kind=None, otype=None, size=543575, hash='XFOV2X6Bw6_zTugTOEBehg', n_files=None, n_observations=None, branch_id=1, created_on_id=1, space_id=1, storage_id=2, run_id=1932, schema_id=None, created_by_id=40, created_at=2026-01-20 19:52:54 UTC, is_locked=False, version_tag=None, is_latest=True)
! returning collection with same hash: Collection(uid='H1AlT19wFq7HdHqZ0000', key='View Habib17 (h5ad) in Vitessce', description=None, hash='hmrSAi0jDsrxvo9Oy3Hiqg', reference=None, reference_type=None, meta_artifact=None, branch_id=1, created_on_id=1, space_id=1, created_by_id=18, run_id=1934, created_at=2026-01-20 20:28:39 UTC, is_locked=False, version_tag=None, is_latest=True); if you intended to query to track this collection as an input, use: ln.Collection.get()
→ returning artifact with same hash: Artifact(uid='JJ58SAXZaCb6Qbna0000', key=None, description='View Habib17 (h5ad) in Vitessce', suffix='.vitessce.json', kind='__lamindb_config__', otype=None, size=1295, hash='yZj12VtmR3rBGZSqmM8iFA', n_files=None, n_observations=None, branch_id=1, created_on_id=1, space_id=1, storage_id=2, run_id=1935, schema_id=None, created_by_id=18, created_at=2026-01-20 20:28:39 UTC, is_locked=False, version_tag=None, is_latest=True); to track this artifact as an input, use: ln.Artifact.get()
→ VitessceConfig: https://lamin.ai/laminlabs/lamindata/artifact/JJ58SAXZaCb6Qbna0000
→ Collection: https://lamin.ai/laminlabs/lamindata/collection/H1AlT19wFq7HdHqZ0000
Note
After running save_vitessce_config, a Vitessce button will appear next to the dataset on the Artifacts or Collections page of the web interface.
If your VitessceConfig object references data from multiple artifacts, the Vitessce button will appear next to a collection that groups these artifacts (on the Collections tab of the Artifacts page).
Note that when using an .h5ad-based artifact, the presence of the corresponding .reference.json file will result in the creation of a collection.
The Vitessce button for this dataset is available on the Collection page.
Visualize an AnnData object (Zarr format)¶
AnnData objects can be saved on-disk to not only .h5ad files, but also to Zarr stores using AnnData’s write_zarr method.
Just like in the above section, we use a Zarr storage that has been previously written with write_zarr() and subset to highly variable genes and ingested into the public laminlabs/lamindata instance in this transform.
adata_zarr_artifact = ln.Artifact.get(key="vitessce_examples/habib17.adata.zarr")
Save a VitessceConfig object¶
You can create a dashboard for one or several datasets by using Vitessce’s component API.
Here, we configure the visualization the same way as above in the .h5ad-based example, with the exception of the ref_artifact parameter, as .zarr-based AnnData objects do not require a Reference Specification for Zarr interoperability.
vc = vit.VitessceConfig(
schema_version="1.0.18",
description=adata_zarr_artifact.description,
)
dataset = vc.add_dataset(name="Habib 2017").add_object(
vit.AnnDataWrapper(
adata_artifact=adata_zarr_artifact,
obs_feature_matrix_path="X",
obs_embedding_paths=["obsm/X_umap"],
obs_embedding_names=["UMAP"],
obs_set_paths=["obs/CellType"],
obs_set_names=["Cell Type"],
)
)
obs_sets = vc.add_view(vit.Component.OBS_SETS, dataset=dataset)
obs_sets_sizes = vc.add_view(vit.Component.OBS_SET_SIZES, dataset=dataset)
scatterplot = vc.add_view(vit.Component.SCATTERPLOT, dataset=dataset, mapping="UMAP")
heatmap = vc.add_view(vit.Component.HEATMAP, dataset=dataset)
genes = vc.add_view(vit.Component.FEATURE_LIST, dataset=dataset)
vc.link_views([scatterplot, heatmap], ["featureValueColormapRange"], [[0.0, 0.1]])
vc.layout(((scatterplot | obs_sets) / heatmap) | (obs_sets_sizes / genes))
Show code cell output
<vitessce.config.VitessceConfig at 0x7fc0a8628f20>
Save the VitessceConfig object.
adata_zarr_vc_artifact = ln.integrations.save_vitessce_config(
vc,
description="View Habib17 in Vitessce",
)
Show code cell output
→ VitessceConfig references these artifacts:
Artifact(uid='Ljx9cSOxELkGisVt0000', key='vitessce_examples/habib17.adata.zarr', description='Habib et al., 2017 Nature Methods, zarr', suffix='.zarr', kind='dataset', otype='AnnData', size=10601062, hash='YPuOwAd5BZ3pbPoGNBH4aw', n_files=208, n_observations=None, branch_id=1, created_on_id=1, space_id=1, storage_id=2, run_id=1932, schema_id=None, created_by_id=40, created_at=2026-01-20 19:53:03 UTC, is_locked=False, version_tag=None, is_latest=True)
→ returning artifact with same hash: Artifact(uid='WLkGgCTmG8eWBzng0000', key=None, description='View Habib17 in Vitessce', suffix='.vitessce.json', kind='__lamindb_config__', otype=None, size=1204, hash='VBBxpSj1tfUHuVKPJXdgKw', n_files=None, n_observations=None, branch_id=1, created_on_id=1, space_id=1, storage_id=2, run_id=1936, schema_id=None, created_by_id=18, created_at=2026-01-20 20:28:40 UTC, is_locked=False, version_tag=None, is_latest=True); to track this artifact as an input, use: ln.Artifact.get()
→ VitessceConfig: https://lamin.ai/laminlabs/lamindata/artifact/WLkGgCTmG8eWBzng0000
→ Dataset: https://lamin.ai/laminlabs/lamindata/artifact/Ljx9cSOxELkGisVt0000
Note
After running save_vitessce_config, a Vitessce button will appear next to the dataset on the Artifacts or Collections page of the web interface.
If your VitessceConfig object references data from multiple artifacts, the Vitessce button will appear next to a Collection that groups these artifacts (on the Collections tab of the Artifacts page).
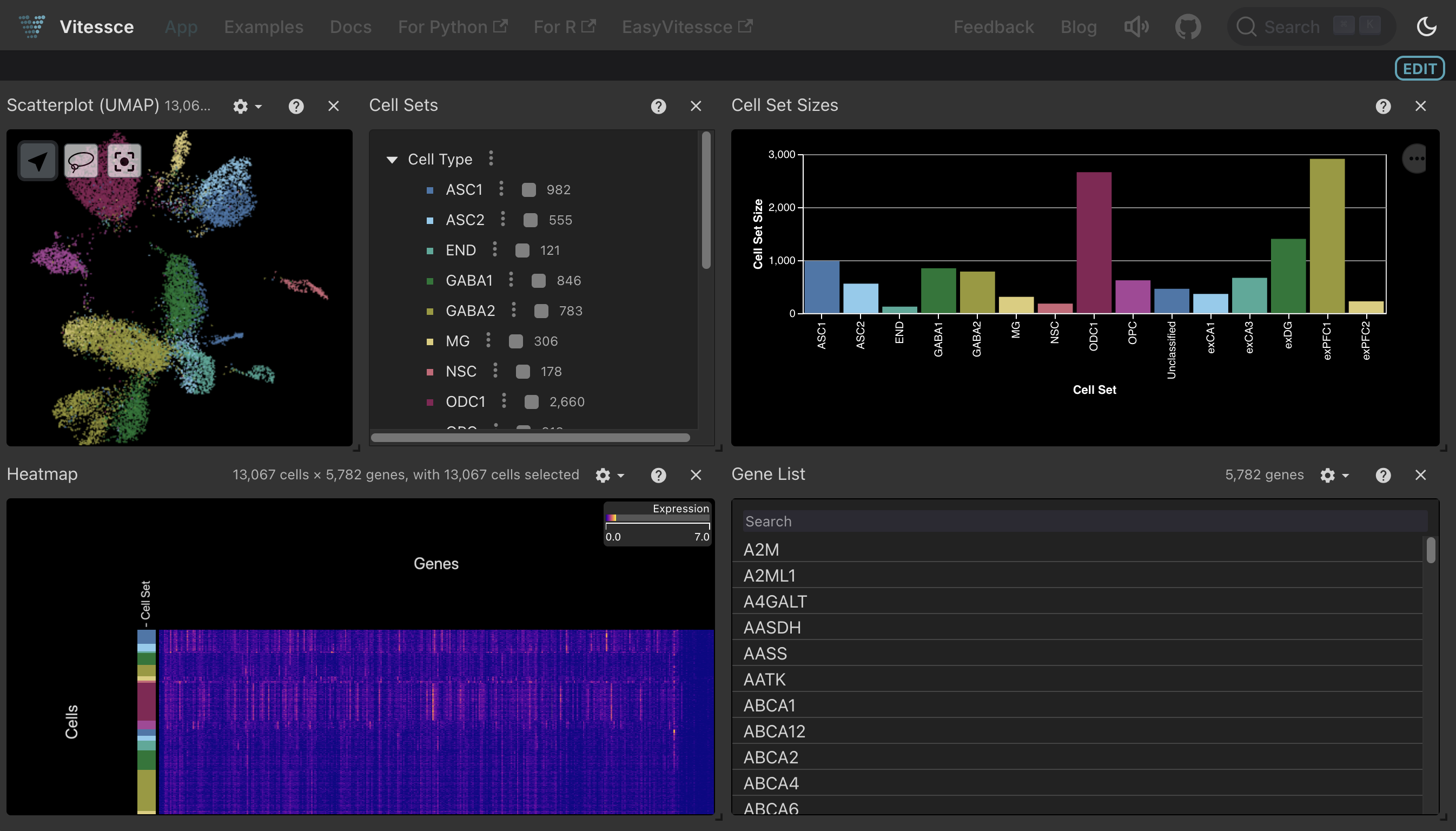
Clicking the Vitessce button for this .zarr-based artifact or .h5ad-based collection launches the same interactive viewer, as both formats represent the same dataset:
Show code cell content
# compare the generated vitessce config to the public one on vitessce/examples (H5AD)
db = ln.DB("vitessce/examples")
public_vc_json = db.Artifact.get("ffUKrGJGNHL3TDhG0000").load()
h5ad_vc_json = h5ad_vc_artifact.load()
assert public_vc_json["layout"] == h5ad_vc_json["layout"]
assert public_vc_json["coordinationSpace"] == h5ad_vc_json["coordinationSpace"]
# compare the generated vitessce config to the public one on vitessce/examples (Zarr)
public_vc_json = db.Artifact.get("J4tMB6qAeHvsgEsp0000").load()
adata_zarr_vc_json = adata_zarr_vc_artifact.load()
assert public_vc_json["layout"] == adata_zarr_vc_json["layout"]
assert public_vc_json["coordinationSpace"] == adata_zarr_vc_json["coordinationSpace"]
→ mapped: Artifact(uid='ffUKrGJGNHL3TDhG0000')
→ mapped: Artifact(uid='J4tMB6qAeHvsgEsp0000')
ln.finish()
Show code cell output
! returning transform with same hash & key: Transform(uid='ZOf7HJEqB4y9000B', key='vitessce.ipynb', description='Vitessce: AnnData [](https://github.com/laminlabs/lamin-spatial/blob/main/docs/vitessce.md)', kind='notebook', hash='X_9z7sCqNkLdG9xAyiHvBA', reference=None, reference_type=None, environment=None, plan=None, branch_id=1, created_on_id=1, space_id=1, created_by_id=18, created_at=2026-02-05 15:28:21 UTC, is_locked=False, version_tag=None, is_latest=False)
• new latest Transform version is: ZOf7HJEqB4y9000B
→ finished Run('seATHQDWnvTX7QIM') after 17s at 2026-05-26 09:52:48 UTC
→ go to: https://lamin.ai/laminlabs/lamindata/transform/ZOf7HJEqB4y9000B
→ to update your notebook from the CLI, run: lamin save /home/runner/work/lamin-spatial/lamin-spatial/docs/vitessce.ipynb