Analyze a collection in memory 
 ¶
¶
Here, we’ll analyze the growing collection by loading it into memory.
This is only possible if it’s not too large.
If your data is large, you’ll likely want to iterate over the collection to train a model, the topic of the next page (
import lamindb as ln
import bionty as bt
ln.track()
Show code cell output
→ connected lamindb: testuser1/test-scrna
→ created Transform('bEtabIli3Ul50000', key='scrna4.ipynb'), started new Run('VnpEWTPONFPbSXtI') at 2026-05-20 14:52:52 UTC
→ notebook imports: bionty==2.4.0 lamindb-core==2.4.2 scanpy==1.12.1
• recommendation: to identify the notebook across renames, pass the uid: ln.track("bEtabIli3Ul5")
collection = ln.Collection.get(key="scrna/collection1")
collection.artifacts.to_dataframe()
Show code cell output
| uid | key | description | suffix | kind | otype | size | hash | n_files | n_observations | ... | is_latest | is_locked | created_at | branch_id | created_on_id | space_id | storage_id | run_id | schema_id | created_by_id | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| id | |||||||||||||||||||||
| 4 | PIreYczLymnjEVDn0000 | scrna/dataset2.h5ad | 10x reference adata, trusted cell type annotation | .h5ad | dataset | AnnData | 855836 | -Finjf36qwZVQLR1AzouyA | None | 70 | ... | True | False | 2026-05-20 14:52:45.408000+00:00 | 1 | 1 | 1 | 1 | 2 | 3 | 1 |
| 1 | GTk9e8L7NkYVSuzT0000 | datasets/conde22.h5ad | None | .h5ad | dataset | AnnData | 57612943 | oHb_G_zCRDhZTJpW_Z5_sm | None | 1648 | ... | True | False | 2026-05-20 14:52:34.154000+00:00 | 1 | 1 | 1 | 1 | 1 | 3 | 1 |
2 rows × 21 columns
If the collection isn’t too large, we can now load it into memory.
Under-the-hood, the AnnData objects are concatenated during loading.
The amount of time this takes depends on a variety of factors.
If it occurs often, one might consider storing a concatenated version of the collection, rather than the individual pieces.
adata = collection.load()
The default is an outer join during concatenation as in pandas:
adata
Show code cell output
AnnData object with n_obs × n_vars = 1718 × 36520
obs: 'donor', 'tissue', 'cell_type', 'assay', 'cell_type_untrusted', 'n_genes', 'percent_mito', 'louvain', 'artifact_uid'
obsm: 'X_umap', 'X_pca'
The AnnData has the reference to the individual artifacts in the .obs annotations:
adata.obs.artifact_uid.cat.categories
Show code cell output
Index(['GTk9e8L7NkYVSuzT0000', 'PIreYczLymnjEVDn0000'], dtype='object')
We can easily obtain ensemble IDs for gene symbols using the look up object:
genes = bt.Gene.lookup(field="symbol")
genes.itm2b.ensembl_gene_id
Show code cell output
'ENSG00000136156'
Let us create a plot:
import scanpy as sc
sc.pp.pca(adata, n_comps=2)
pca_itm2b_fig = sc.pl.pca(
adata,
color=genes.itm2b.ensembl_gene_id,
title=(
f"{genes.itm2b.symbol} / {genes.itm2b.ensembl_gene_id} /"
f" {genes.itm2b.description}"
),
return_fig=True
)
pca_itm2b_fig.figure.savefig('pca_itm2b.pdf')
Show code cell output
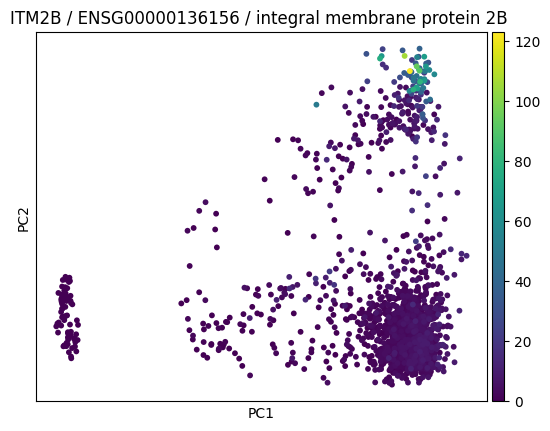
We could save a plot as a pdf and then see it in the flow diagram:
artifact = ln.Artifact(
"pca_itm2b.pdf", description="My result on ITM2B"
).save()
artifact.view_lineage()
Show code cell output
But given the image is part of the notebook, we can also rely on the report that we create when saving the notebook:
ln.finish()